Authors:
Yuan Liu1 and Zhaoxia Cui1,2*
1Key Laboratory of Experimental Marine Biology, Institute of Oceanology, Chinese Academy of Sciences, Qingdao, 266071, China
2National & Local Joint Engineering Laboratory of Ecological Mariculture, Qingdao, 266071, China
Received: 05 March, 2014; Accepted: 04 April, 2015; Published: 06 April, 2015
Zhaoxia Cui, Key Laboratory of Experimental Marine Biology, Institute of Oceanology, Chinese Academy of Sciences, 7 Nanhai Road, Qingdao 266071, China; Tel: +86 532 82898509; Fax: +86 532 82898509; Email:
Liu Y, Cui Z (2015) Molecular Cloning and Characterization of Dmc1 from the Chinese Mitten Crab (Eriocheir sinensis). Int J Aquac Fishery Sci 1(1): 024-029. DOI: 10.17352/2455-8400.000005
© 2015 Liu Y, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Eriocheir sinensis; Dmc1; Meiosis; Expression analysis
Dmc1, a member of the RecA/Rad51 superfamily, is essential for meiotic recombination. In this study, a Dmc1 gene (EsDmc1) was identified from screening the larval transcriptomes of Chinese mitten crab Eriocheir sinensis. The full-length cDNA of EsDmc1 was 1478 bp long and contained a 1026 bp open-reading frame encoding 341 amino acids. The genomic fragment of EsDmc1 contained two exons separated by one intron. Several tandem repeats were found in intron. The deduced EsDmc1 protein contained motifs conserved in the RacA/Rad51 superfamily, including the Walker A and B motifs, and L1 and L2 loops. EsDmcl shared 87.3%, 86.5% and 77.4% identity with its homologues in Litopenaeus vannamei, Penaeus monodon and Ixodes scapularis, respectively. Phylogenetic analysis revealed that EsDmc1 had a closer relationship with Dmc1s from arthropods than vertebrates. The EsDmcl transcripts could be detected in all examined larval stages with the highest expression level in the fifth zoeal stage. These results suggest that EsDmc1 could be expressed before reproductive maturity and might have complex functions in crab reproduction.
Introduction
The Chinese mitten crab Eriocheir sinensis (Henri Milne Edwards 1854) is a commercially important species in Southeast Asia and has been widely farmed in ponds, reservoirs and lakes of China since 1990s [1]. Like other crabs, the mitten crabs exhibit bimodal growth patterns in which males exhibit higher growth rate and larger size than females [2-4]. However, female mitten crabs have greater economic value than males. In addition, sexual precocity has occurred in cultured E. sinensis populations [5] and caused catastrophic economic losses to crab aquaculture. Understanding the mechanisms involved in reproduction or sexual development and characterization of the related genes in E. sinensis would be helpful for the aquaculture industry.
Dmc1 (disrupted meiotic cDNA), a member of the RecA/Rad51 superfamily that encodes a DNA recombinase, plays a central role in meiotic recombination [6,7]. The Dmc1 gene was initially identified in budding yeast Saccharomyces cerevisiae as a meiosis-specific gene [8]. Afterwards, Dmc1 genes have been isolated from higher eukaryotes, such as human and mouse [9]. In mammals, Dmc1 was also a specifically expressed gene in testicular germ cells and embryonic ovaries in the period of meiosis. Further studies showed that Dmc1 mutations cause meiotic arrest at the zygotene stage without homologue synapsis [10] or with an occasional synapsis between non-homologues [11]. All of these suggest the crucial role of Dmc1 during meiosis appears to be conserved from yeast to mammals.
Recently, Dmc1 genes have been identified and characterized from several aquatic animals, such as Japanese eel [12], cyprinid fishes [13], and only two crustacean species, giant tiger shrimp Penaeus monodon [14,15] and whiteleg shrimp Litopenaeus vannamei [16]. Despite the finding of Dmc1 from testis transcriptome [17], no study has described the sequence characterization and expression pattern of Dmc1 in E. sinensis. Here, a Dmc1 gene (designated as EsDmc1) showing a predicted amino acid sequence similarity to L. vannamei was found from screening the transcriptomes of larval E. sinensis. The main objectives of the present study are to clone this gene from E. sinensis, detect genomic organization and investigate the expression pattern during larval developmental stages based on transcriptome data.
Materials and Methods
cDNA library construction and gene annotation
Four libraries were conducted from whole bodies of E. sinensis at four larval developmental stages, including the fifth zoeal stage (Z5), the megalopa stages before and after desalination (MB and MA), and the first juvenile instar (J1). Transcriptome sequencing was carried on an Illumina HiSeqTM 2000 platform. A total of 23.48 Gb clean data were generated from larval transcriptomes [18-20]. BLASTx analysis revealed that a unigene (comp19040_c0_seq1) was homologous to Dmc1 in L. vannamei (ADM45305).
Full-length cDNA sequence determination
Gene specific primer (P1), nested primer (P2) and oligo (dT)-adaptor (Table 1) were used to clone the 3' end of EsDmc1cDNA by 3' RACE technique. Initial amplification was carried out with P1 and oligo (dT)-adaptor, followed by nested PCR using P2 and oligo (dT)-adaptor. The amplification was performed in a 25 µl reaction volume containing 17.3 µl sterile distilled H2O, 2.5 µl of 10×PCR buffer, 1.5 µl of MgCl2 (25 mM), 0.5 µl of dNTP (10 mM), 1 µl of each primer (5 µM), 0.2 µl (1 U) of Taq polymerase (TaKaRa), and 1 µl of template. The PCR reactions were performed on TaKaRa PCR Thermal Cycler Dice Model TP600 (Takara Bio Inc.) with an initial denaturation at 94°C for 3 min, followed by 35 cycles of denaturation at 94°C for 30 s, annealing at 56°C for 50 s, extension at 72°C for 1 min, and a final extension at 72°C for 10 min. The PCR products were gel-purified and cloned into pMD18-T simple vector (TaKaRa). After being transformed into the competent cells of Escherichia coli DH5α, the positive recombinants were identified through anti-Amp selection and PCR screening with M13-47 and RV-M primers (Table 1). Three of the positive clones were sequenced to verify the full-length cDNA of EsDmc1.
Genomic DNA amplification
Genomic DNA was extracted from the muscle tissue by standard phenol-chloroform method [21]. To detect the genomic structure of EsDmc1, two gene-specific primers (P3 and P4, see Table 1) were designed according to the obtained cDNA sequence. The PCR was performed in a 25 µl reaction volume containing 17.3 µl sterile distilled H2O, 2.5 µl of 10×PCR buffer, 1.5 µl of MgCl2 (25 mM), 0.5 µl of dNTP (10 mM), 1 µl of each primer (5 µM), 0.2 µl (1 U) of Taq polymerase (TaKaRa), and 1 µl of DNA template (approximately 30 ng). The PCR temperature profile was 94°C for 3 min, followed by 34 cycles of 94°C for 30 s, 55°C for 50 s, 72°C for 1 min, and a final extension at 72°C for 10 min. The PCR products were cloned and sequenced according to the method described above.
Sequence and phylogenetic analysis
Blast algorithm at National Center for Biotechnology Information (http://www.ncbi.nlm.gov/blast/) was used to search the homology of nucleotide and protein sequences. The deduced amino acid sequence was analyzed with the Expert Protein Analysis System (http://www.expasy.org/). SignalP 3.0 program was utilized to predict the presence and location of signal peptide, and the cleavage sites in amino acid sequences (http://www.cbs.dtu.dk/services/SignalP).
Multiple amino acid sequence alignment was performed using the Clustal X with the default settings [22]. A neighbor-joining (NJ) tree with bootstrap values was constructed for phylogenetic analysis using MEGA 4.0 [23]. All the reference sequences for phylogenetic analysis were derived from GenBank.
Expression analyses based on transcriptome data
The reads for EsDmc1 were counted by mapping reads to assembled unigene sequences in the four larval transcriptomes. The expression of EsDmc1 was calculated using the reads per kb per million reads (RPKM) method [24]. The fold change values > 2 and false discovery rates (FDR) adjusted significance values < 0.05 (-log10 (0.05) = 1.3) were considered as the threshold to judge the significance of expression.
Results and Discussion
cDNA cloning and sequence analysis of EsDmc1
The full-length cDNA sequence of EsDmc1 was obtained by overlapping the corresponding unigene with the amplified fragments. It was 1478 bp, and consisted of a 5'-UTR of 213 bp, 3'-UTR of 239 bp and an open-reading frame (ORF) of 1026 bp. A canonical polyadenylation signal-sequence (AATAAA) and poly (A) tail was detected in EsDmc1. The ORF encoded 341 amino acids and no signal peptide was identified (Figure 1). The estimated molecular weight of EsDmc1 was 37.49 kDa and its theoretical isoelectric point was 5.41. The sequence of EsDmc1 was deposited in GenBank under the accession number KP876487.
Genomic organization of EsDmc1
Until now, no genomic sequence of Dmc1 has been identified in crustaceans. The amplified genomic DNA fragment of EsDmc1 was 856 bp and deposited in GenBank under accession number KP876488. By aligning with the corresponding cDNA sequence, the exon-intron boundaries of EsDmc1 were determined (Figure 2). The EsDmc1 genomic sequence consisted of two exons (248 and 355 bp) separated by one intron (253 bp). All splice sites in EsDmc1 followed the canonical GT/AG splicing recognition rule. Several mononucleotide repeats (T)4-7 and one pure dinucleotide repeat (GC)3 were found in the intron (Figure 2). Intronic microsatellites found in EsDmc1 will be helpful in understanding the genetic structure of Dmc1s and studying the possible roles of microsatellites.
-

Figure 1:
Nucleotide and deduced amino acid sequences of EsDmc1 from Eriocheir sinensis. Numbers on the right of the sequence give the positions of the last nucleotide and amino acid on each line, respectively. The start and stop codons are boxed and the classical polyadenylation signal in the 3′-UTR is underlined.
-

Figure 2:
Genomic organization (A) and nucleotide sequence (B) of EsDmc1. The exon and intron sequences are illustrated with capital and lowercase letters, respectively. The tandem repeats in intron are shadowed. Intron dinucleotide acceptor and donor sites (gt/ag) for RNA splicing are underlined.
Homologous and phylogenetic analysis of EsDmc1
BLAST analysis indicated that the deduced amino acid sequence of EsDmc1 shared significant identity with other reported Dmc1s. EsDmc1 displayed 87.3% identity with published Dmc1 of L. vannamei (LvDmc1), 86.5% with P. monodon (PmDmc1), 77.4% with Ixodes scapularis (IsDmc1), 75.9% with Anguilla japonica (AjDmc1) and 74.8% with Homo sapiens (HsDmc1). As reported in LvDmc1 [16], multiple alignments revealed that EsDmc1 contained motifs conserved in the RecA/Rad51 superfamily, including L1 and L2 loops, which are single-strand DNA binding motifs [25,26], and the Walker A and B motifs of the ATP binding site [26,27] (Figure 3). Moreover, the helix-hairpin-helix (HhH) motif, a double-strand DNA binding site that is commonly present in Rad51 and Dmc1, but not in RecA [26,28,29], was also detected in EsDmc1.
-

Figure 3:
Multiple alignments of EsDmc1 with other known Dmc1s. Amino acid residues with 100% conservation are shaded in dark, and similar amino acids are shaded in grey. Numbers on the right indicate the amino acid position of the different sequences. Each domain conserved in the RacA/Rad51 superfamily is underlined. The species and the GenBank accession numbers are as follow: Anguilla japonica AjDmc1 (BAD98459), Danio rerio DrDmc1 (BAD98462), Homo sapiens HsDmc1 (NP_008999), Litopenaeus vannamei LvDmc1 (ADM45305), Penaeus monodon PmDmc1 (ACC62173), Ixodes scapularis IsDmc1 (XP_002436011) and the budding yeast Saccharomyces cerevisiae ScDmc1 (NP_011106).
Furthermore, a phylogenetic tree was constructed based on 14 amino acid sequences of Dmc1 members by NJ method (Figure 4). The Clustal X alignment consisted of 346 characters, 149 being constant, 195 variable but parsimony-uninformative and 103 informative. The phylogenetic tree showed that the Dmc1s formed two major clades: one with four arthropods and the other containing two subclades from the vertebrates. EsDmc1 was clustered together with Dmc1s from the whiteleg shrimp L. vannamei and the giant tiger shrimp P. monodon, and then had a closer relationship with blacklegged tick I. scapularis. Both homology and phylogenetic analysis together showed that EsDmc1 is a clearly member of Dmc1 homologous belonging to RecA/Rad51 superfamily.
-
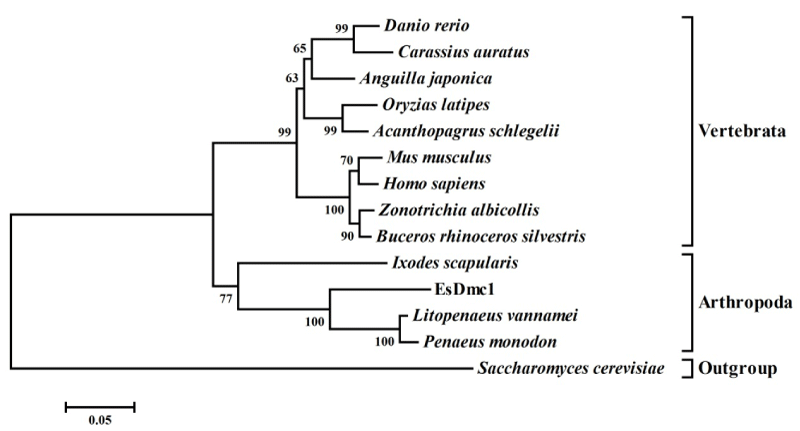
Figure 4:
A bootstrapped NJ tree inferred from amino acid sequences of EsDmc1 and other reported Dmc1s. The species and the GenBank accession numbers are as follow: Danio rerio DrDmc1 (BAD98462), Carassius auratus (ABQ23182), Anguilla japonica AjDmc1 (BAD98459), Oryzias latipes (ACH91672), Acanthopagrus schlegelii (AFV31617), Mus musculus (NP_034189), Homo sapiens HsDmc1 (NP_008999), Zonotrichia albicollis (XP_005488680), Buceros rhinoceros silvestris (XP_010136730), Ixodes scapularis IsDmc1 (XP_002436011), Litopenaeus vannamei LvDmc1 (ADM45305), and Penaeus monodon PmDmc1 (ACC62173). The budding yeast Saccharomyces cerevisiae ScDmc1 (NP_011106) serves as outgroups. Numbers in each branch indicate bootstrap values.
Expression of EsDmc1 in larval developmental stages
Most studies on expression profiles of Dmc1 transcripts have focused on the gonads and somatic tissues of adults or subadults [12,14-16]. However, little is known about the expression pattern of Dmc1 in larval developmental stages. In the present study, the transcripts of EsDmc1 could be detected in all examined larval stages (Figure 5). EsDmc1 had the highest expression level in Z5, moderate expression in MB and MA, while only a week expression level in JI. Moreover, the expression of EsDmc1 in Z5 was significantly higher than that in MA and JI. This expression pattern would suggest EsDmc1 could be expressed at earlier developmental stages before reproductive maturity in E. sinensis. Contrary to the results from L. vannamei, P. monodon Dmc1 did not show gonad-specific expression [14-16], indicating the complex roles of Dmc1 in crustaceans. Further investigations regarding crab Dmc1, particularly functional studies, are needed to determine the biological roles of this gene in crustaceans.
-
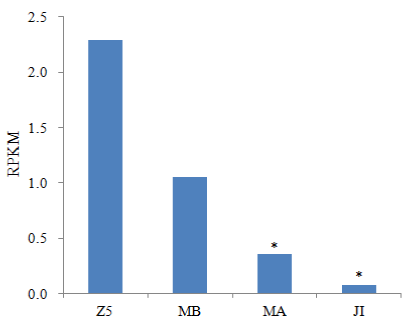
Figure 5:
Expression levels of EmDmc1 transcripts in four larval transcriptomes of Eriocheir sinensis. Significant differences across Z5 are indicated with an asterisk.
Conclusion
This study reported a Dmc1 gene in the Chinese mitten crab, E. sinensis, encoding amino acid sequences characteristic of the RecA/Rad51 superfamily. Multiple alignment and phylogenetic analysis showed EsDmc1 was a member of Dmc1 homologues. Different from the previous studies in other species, EsDmc1 transcripts were found to be expressed at earlier larval stages, from Z5 to JI. Our study provides more information about crustacean Dmc1, and will facilitate further investigations of the molecular mechanisms involved in crab reproduction.
Acknowledgement
This research was supported by grants from Chinese National '863' Project (No. 2012AA10A409), National Natural Science Foundation of China (41206147) and Open Fund of Zhejiang Provincial Top Key Discipline of Aquaculture (xkzsc02).
-
-
- Chen D, Zhang M, Shrestha S (2007) Compositional characteristics and nutritional quality of Chinese mitten crab (Eriocheir sinensis). Food Chem 103: 1343-1349.
- Hartnoll RG, Growth (1985) In: Bliss DE, editor. The Biology of Crustacea. New York: Academic Press. 111-197.
- Lee HH, Hsu CC (2003) Population biology of the swimming crab Portunus sanguinolentus in the waters off Northern Taiwan. J Crustacean Biol 23: 691-699.
- Mokhtari M, Savari A, Rezai H, Kochanian P, Bitaab A (2008) Population ecology of fiddler crab, Uca lactea annulipes (Decapoda: Ocypodidae) in Sirik mangrove estuary, Iran. Estuar Coast Shelf S 76: 273-281.
- Rudnick DA, Hieb K, Grimmer KF, Resh VH (2003) Patterns and processes of biological invasion: the Chinese mitten crab in San Francisco Bay. Basic Appl Ecol 4: 249-262.
- Shinohara A, Shinohara M (2004) Roles of RecA homologues Rad51 and Dmc1 during meiotic recombination. Cytogenet Genome Res 107: 201-207.
- Kagawa W, Kurumizaka H (2010) From meiosis to postmeiotic events: uncovering the molecular roles of the meiosis-specific recombinase Dmc1. FEBS J 277: 590-598.
- Bishop DK, Park D, Xu L, Kleckner N (1992) DMC1: a meiosisspecific yeast homolog of E. coli recA required for recombination, synaptonemal complex formation, and cell cycle progression. Cell 69: 439-456.
- Habu T, Taki T, West A, Nishimune Y, Morita T (1996) The mouse and human homologs of DMC1, the yeast meiosis-specific homologous recombination gene, have a common unique form of exon-skipped transcript in meiosis. Nucleic Acid Res 24: 470-477.
- Pittman DL, Cobb J, Schimenti KJ, Wilson LA, Cooper DM, et al. (1998) Meiotic prophase arrest with failure of chromosome synapses in mice deficient for Dmc1, a germline-specific RecA homolog. Mol Cell 1: 697-705.
- Yoshida K, Kondoh G, Matsuda Y, Habu T, Nishimune Y, et al. (1998) The mouse RecA-like gene Dmc1 is required for homologous chromosome synapsis during meiosis. Mol Cell 1: 707-718.
- Kajiura-Kobayashi H, Kobayashi T, Nagahama Y (2005) Cloning of cDNAs and the differential expression of A-type cyclins and Dmc1 during spermatogenesis in the Japanese eel, a teleost fish. Dev Dyn 232: 1115-1123.
- Tao M, Liu S, Long Y, Zeng C, Liu J, et al. (2008) The cloning of Dmc1 cDNAs and a comparative study of its expression in different ploidy cyprinid fishes. Sci China C Life Sci 51: 38-46.
- Leelatanawit R, Klinbunga S, Aoki T, Hirono I, Valyasevi R, et al. (2008) Suppression subtractive hybridization (SSH) for isolation and characterization of genes related to testicular development in the giant tiger shrimp Penaeus monodon. BMB Rep 41: 796-802.
- Wongsurawat T, Leelatanawit R, Thamniemdee N, Uawisetwathana U, Karoonuthaisiri N, et al. (2010) Identification of testis-relevant genes using in silico analysis from testis ESTs and cDNA microarray in the black tiger shrimp (Penaeus monodon). BMC Mol Biol 11: 55.
- Okutsu T, Kang BJ, Miwa M, Yoshizaki G, Maeno Y, et al. (2010) Molecular cloning and characterization of Dmc1, a gene involved in gametogenesis, from the whiteleg shrimp Litopenaeus vannamei. Fish Sci 76: 961-969.
- Zhang W, Wan H, Jiang H, Zhao Y, Zhang X, et al. (2011) A transcriptome analysis of mitten crab testes (Eriocheir sinensis). Genet and Mol Biol 34: 136-141.
- Li Y, Hui M, Cui Z, Liu Y, Song C, et al. (2015) Comparative transcriptomic analysis provides insights into the molecular basis of the metamorphosis and nutrition metabolism change from zoeae to megalopae in Eriocheir sinensis. Comp Biochem Phys D 13: 1-9.
- Hui M, Liu Y, Song C, Li Y, Shi G, et al. (2014) Transcriptome changes in Eriocheir sinensis megalopae after desalination provide insights into osmoregulation and stress adaption in larvae. PloS one 9: e114187.
- Song C, Cui Z, Hui M, Liu Y, Li Y, et al. (2015) Comparative transcriptomic analysis provides insights into the molecular basis of brachyurization and adaptation to benthic lifestyle in Eriocheir sinensis. Gene 558: 88-98.
- Sambrook J, Russell DW (2002) Molecular cloning: a laboratory manual. New York: Cold Spring Harbor Laboratory Press.
- Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25: 4876-4882.
- Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Bio Evol 24: 1596-1599.
- Mortazavi A, Williams BA, Mccue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5: 621-628.
- Story RM, Steitz TA (1992) Structure of the recA protein-ADP complex. Nature 355:375-376.
- Chen LT, Ko TP, Chang YC, Lin KA, Chang CS, et al. (2007) Crystal structure of the left-handed archaeal RadA helical filament: identification of a functional motif for controlling quaternary structures and enzymatic functions of RecA family proteins. Nucleic Acids Res 35: 1787-1801.
- Hong EL, Shinohara A, Bishop D (2001) Saccharomyces cerevisiae Dmc1 protein promotes renaturation of single-strand DNA (ssDNA) and assimilation of ssDNA into homologous supercoiled duplex DNA. J Biol Chem 276: 41906-41912.
- Aihara H, Ito Y, Kurumizaka H, Yokoyama S, Shibata T () The N-terminal domain of the human Rad51 protein binds DNA: structure and a DNA binding surface as revealed by NMR. J Mol Biol 290: 495-504.
- Kinebuchi T, Kagawa W, Kurumizaka H, Yokoyama S (2005) Role of the N-terminal domain of the human DMC1 protein in octamer formation and DNA binding. J Biol Chem 280: 28382-28387.
-
-









Table 1:
Primers used in this study.
Table 1: Primers used in this study.